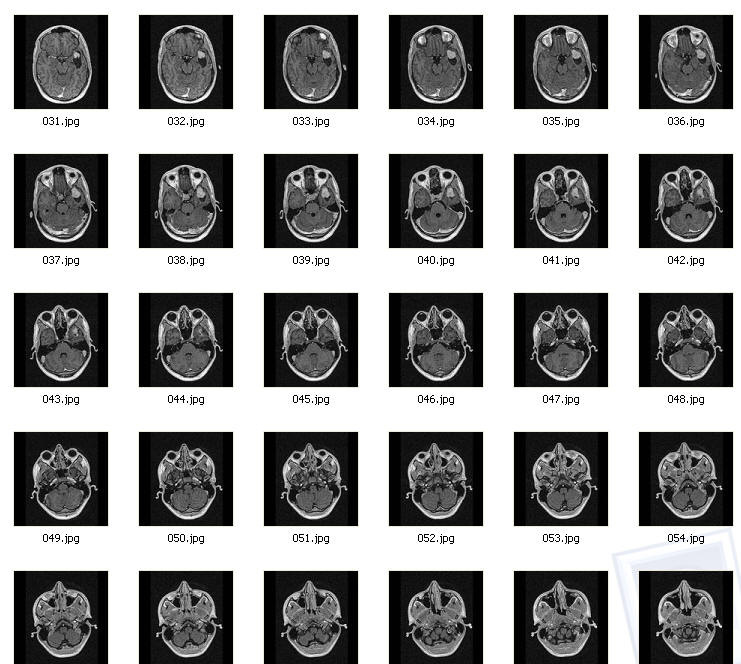
DICOM files
DICOM file
clc;
close all;
clear all;
% N=input('Enter number of images');
k=1;
% N=5;
N=5;
for ic=1:1:N;
A='C:\Data\January15\31\';
B=num2str(ic);
C='.jpg';
path=strcat(A,B,C)
I=imread(path);
[m,n,q]=size(I);
figure()
imshow(I);
if(q==3)
I=rgb2gray(I);
end
figure()
imshow(I);
I=edge(I,'sobel');
figure()
imshow(I);
for i=1:1:m
for j=1:1:n
if(I(i,j)==1)
x(k)=m-i;
y(k)=j;
z(k)= ic;
k=k+1;
end
end
end
figure(24)
hold all
plot3(y,x,z,'*');
end
close all;
clear all;
% N=input('Enter number of images');
k=1;
% N=5;
N=5;
for ic=1:1:N;
A='C:\Data\January15\31\';
B=num2str(ic);
C='.jpg';
path=strcat(A,B,C)
I=imread(path);
[m,n,q]=size(I);
figure()
imshow(I);
if(q==3)
I=rgb2gray(I);
end
figure()
imshow(I);
I=edge(I,'sobel');
figure()
imshow(I);
for i=1:1:m
for j=1:1:n
if(I(i,j)==1)
x(k)=m-i;
y(k)=j;
z(k)= ic;
k=k+1;
end
end
end
figure(24)
hold all
plot3(y,x,z,'*');
end
Video
Papers
| braintumer.zip | |
| File Size: | 5576 kb |
| File Type: | zip |
| braintumer.z01 | |
| File Size: | 8388 kb |
| File Type: | z01 |
| braintumer.z02 | |
| File Size: | 8388 kb |
| File Type: | z02 |
Blackbooks
Thesis 1
Thesis 2
Thesis 3
Thesis 4
Thesis 5
Thesis 6
Thesis 7
Thesis 8
Thesis 9
Thesis 10
Thesis 11
Thesis 12
Thesis 13
Thesis 2
Thesis 3
Thesis 4
Thesis 5
Thesis 6
Thesis 7
Thesis 8
Thesis 9
Thesis 10
Thesis 11
Thesis 12
Thesis 13
Presentations
| braintumor.pptx | |
| File Size: | 1013 kb |
| File Type: | pptx |
Block diagram
Flow chart
clear all %no variables
close all %no figures
clc %empty command window
prefix = 'C:\Data\February14\22\MRI Brain Scan\Series 8\I0000'; %file name prefix
fnum = 417:476; %file numbers
ext = '_anon.dcm'; %extension
fname = [prefix num2str(fnum(1)) ext]; % complete file name
info = dicominfo(fname) % extract information
voxel_size = [info.PixelSpacing; info.SliceThickness]' % zstack infor
hWaitBar = waitbar(0,'Reading DICOM files'); % user waitwindow
for i=length(fnum):-1:1 %read all the image
fname = [prefix num2str(fnum(i)) ext]; %changing file name
D(:,:,i) = uint16(dicomread(fname));
waitbar((length(fnum)-i)/length(fnum))
end
delete(hWaitBar)
% for i=length(fnum):-1:1
% figure()
% imshow(uint8(D(:,:,i)))
% end
whos D % confrims D is presents
i = 30; %middle slice user decide
im = squeeze(D(:,:,i)); % preffered existance of brain
max_level = double(max(D(:))); %detect max
imtool(im,[0 max_level]) %tool allows us to viwe all the points and corrosponding grayscal image
fig1 = figure;% covert from imtool to figure window
max_level = double(max(D(:))); %determine maximum level
imshow(im,[0 max_level]) % displays maximum level normalized to 255
title('Coronal Slice #30')
set(fig1,'position',[601 58 392 314])
imtool close all
colorbar
colormap jet
%
% %3D visualization (doc: contourslice, isosurface & isocap)
% docsearch('visualizing mri data')
%
% %explore 3D volumetric data using Slice-O-Matic GUI tool (nobkpt)
% addpath('D:\work\Demos\others\sliceomatic')
% sliceomatic(double(D))
% %ref: submission #780 @ www.mathworks.com/matlabcentral (nobkpt)
% hSlico1 = gcf;
% daspect(1./voxel_size)
% movegui('northwest')
%
% %reorient data for easier interpretation (stand patient up)
% D = permute(D,[3 2 1]);
% voxel_size = voxel_size([1 3 2]);
% for i=1:3
% D = flipdim(D,i);
% end
% whos D
%
% %explore rotated 3D volume (new Slice-O-Matic viwer) - nobkpt
% if ishandle(hSlico1), delete(hSlico1), end
% sliceomatic(double(D))
% daspect(1./voxel_size)
% hSlico2 = gcf;
% set(hSlico2,'position',[455 63 560 420])
%
% %intensity distribution also useful (more custom graphics)
% %max_level = double(max(D(:)));
% my_map = jet(max_level);
% fig2 = figure;
%
% %intensity distribution - top 2/3 (nobkpt)
% subplot(3,1,1:2)
% hist(double(im(:)),max_level)
% axis([0 max_level 0 900])
% title('Distribution')
%
% %color scale - bottom 1/3 (nobkpt)
% subplot(3,1,3)
% imagesc(1:max_level)
% colormap(my_map)
% xlim([0 max_level])
% set(gca,'ytick',[])
% ylabel('Color Map')
% xlabel('Intensity')
% set(fig2,'position',[22 60 560 300],'render','zbuffer')
% set(fig1,'position',[601 68 392 314])
% figure(fig1)
% %% Segmentation
% %----------------------------------------------------------------------
%
% %ignore low levels (backround air, CSF & other soft? tissues)
% %using custom GUI tool to select best threshold level
% im = imrotate(squeeze(D(30,:,:)),90);
% figure(hSlico2)
%
% %remove some figures (no longer needed) - nobkpt
% if ishandle(fig1), delete(fig1), end
% if ishandle(fig2), delete(fig2), end
% doc graythresh
%
% %custom GUI tool (nobkpt)
% thresh_tool(im)
%
% %duplicate original data set for later reference (nobkpt)
% D1 = D;
%
% %apply some thresholding rules to ignore certain parts of data
% D(D<=40) = 0; %ignore low levels (CSF & air)
% D(D>=100) = 0; %ignore high levels (skull & other hard? tissues)
% D(:,:,1:60) = 0; %ignore spatially low positions (below brain mass)
% update_SliceOMatic(double(D),hSlico2)
%
% %erode away thick layer (dissolve thin surrounding tissues)
% blk = ones([3 7 7]);
% D = imerode(D,blk);
% update_SliceOMatic(double(D),hSlico2)
%
% %isolate brain mass (bwlabeln)
% doc bwlabel
% lev = graythresh(double(im)/max_level) * max_level;
% bw = (D>=lev);
% L = bwlabeln(bw);
%
% %connected region properties - how many, how big?
% doc regionprops
% stats = regionprops(L,'Area')
% A = [stats.Area];
% biggest = find(A==max(A))
%
% %remove smaller scraps
% D(L~=biggest) = 0;
% update_SliceOMatic(double(D),hSlico2)
%
% %grow back main region (brian mass) - nobkpt
% D = imdilate(D,blk);
% update_SliceOMatic(double(D),hSlico2)
%
% %separate white vs. gray matter
% im = imrotate(squeeze(D(30,:,:)),90);
% figure(hSlico2)
% lev2 = thresh_tool(im,'gray')
%
% %partition brain mass (nobkpt)
% lev2 = 67;
% L = zeros(size(D)); %0=outside brain (head/air)
% L(D<lev2 & D>0) = 2; %2=gray matter
% L(D>=lev2) = 3; %3=white matter
%
% %new Slice-O-Matic viewer (label matrix) - nobkpt
% sliceomatic(L)
% hSlico3 = gcf;
% daspect(1./voxel_size)
% set(hSlico3,'position',[455 63 560 420])
%
% %remove previous slicomatic viewer (nobkpt)
% if ishandle(hSlico2), delete(hSlico2), end
%
% %% Volumetric Measurements (voxel counting)
% %----------------------------------------------------------------------
%
% %total volume of brain (liters)
% brain_voxels = length(find(L(:)>1));
% brain_volume = brain_voxels*prod(voxel_size)/1e6
%
% %volume of gray matter (liters) - nobkpt
% gray_voxels = length(find(L(:)==2));
% gray_volume = gray_voxels*prod(voxel_size)/1e6
%
% %volume of white matter (liters) - nobkpt
% white_voxels = length(find(L(:)==3));
% white_volume = white_voxels*prod(voxel_size)/1e6
%
% %density calculations (volume ratios) - nobkpt
% gray_fraction = gray_volume/brain_volume
% white_fraction = white_volume/brain_volume
%
% return
%
% %% Separate head from background for visualization (advanced maneuver)
% %----------------------------------------------------------------------
%
% %duplicate data
% L1 = L;
%
% %exterior of head (connected inside through ears)
% %new Slice-O-Matic viewer (binary data) - nobkpt
% BW = (D1<lev1);
% %sliceomatic(double(BW))
% %hSlico3 = gcf;
% %daspect(1./voxel_size)
% %set(hSlico3,'position',[455 63 560 420])
%
% %melt away layer (separate small interior volumes) - nobkpt
% BW = imerode(BW,blk);
% %update_SliceOMatic(double(BW),hSlico3)
%
% %how many connected regions? (nobkpt)
% L = bwlabeln(BW);
% stats = regionprops(L,'Area')
%
% %which one biggest? (nobkpt)
% A = [stats.Area];
% biggest = find(A==max(A))
% BW(L~=biggest) = 0;
% %update_SliceOMatic(double(BW),hSlico3)
%
% %grow layer back (nobkpt)
% BW = imdilate(BW,blk);
%
% %label head voxels (different from air) - nobkpt
% L = L1;
% L(BW<1 & L1<2) = 1; %1=head (0=air, 2=gray, 3=white)
%
% %update final display (nobkpt)
% update_SliceOMatic(L,hSlico3)
%
% %% additional volumetric measurements
% %----------------------------------------------------------------------
%
% %volume ratio occupied by patient (nobkpt)
% scan_voxels = numel(L);
% head_voxels = length(find(L(:)>0));
% scan_density = head_voxels/scan_voxels
%
% %volume ratio of brain in head (nobkpt)
% brain_voxels = length(find(L(:)>1));
% brain_density = brain_voxels/head_voxels
close all %no figures
clc %empty command window
prefix = 'C:\Data\February14\22\MRI Brain Scan\Series 8\I0000'; %file name prefix
fnum = 417:476; %file numbers
ext = '_anon.dcm'; %extension
fname = [prefix num2str(fnum(1)) ext]; % complete file name
info = dicominfo(fname) % extract information
voxel_size = [info.PixelSpacing; info.SliceThickness]' % zstack infor
hWaitBar = waitbar(0,'Reading DICOM files'); % user waitwindow
for i=length(fnum):-1:1 %read all the image
fname = [prefix num2str(fnum(i)) ext]; %changing file name
D(:,:,i) = uint16(dicomread(fname));
waitbar((length(fnum)-i)/length(fnum))
end
delete(hWaitBar)
% for i=length(fnum):-1:1
% figure()
% imshow(uint8(D(:,:,i)))
% end
whos D % confrims D is presents
i = 30; %middle slice user decide
im = squeeze(D(:,:,i)); % preffered existance of brain
max_level = double(max(D(:))); %detect max
imtool(im,[0 max_level]) %tool allows us to viwe all the points and corrosponding grayscal image
fig1 = figure;% covert from imtool to figure window
max_level = double(max(D(:))); %determine maximum level
imshow(im,[0 max_level]) % displays maximum level normalized to 255
title('Coronal Slice #30')
set(fig1,'position',[601 58 392 314])
imtool close all
colorbar
colormap jet
%
% %3D visualization (doc: contourslice, isosurface & isocap)
% docsearch('visualizing mri data')
%
% %explore 3D volumetric data using Slice-O-Matic GUI tool (nobkpt)
% addpath('D:\work\Demos\others\sliceomatic')
% sliceomatic(double(D))
% %ref: submission #780 @ www.mathworks.com/matlabcentral (nobkpt)
% hSlico1 = gcf;
% daspect(1./voxel_size)
% movegui('northwest')
%
% %reorient data for easier interpretation (stand patient up)
% D = permute(D,[3 2 1]);
% voxel_size = voxel_size([1 3 2]);
% for i=1:3
% D = flipdim(D,i);
% end
% whos D
%
% %explore rotated 3D volume (new Slice-O-Matic viwer) - nobkpt
% if ishandle(hSlico1), delete(hSlico1), end
% sliceomatic(double(D))
% daspect(1./voxel_size)
% hSlico2 = gcf;
% set(hSlico2,'position',[455 63 560 420])
%
% %intensity distribution also useful (more custom graphics)
% %max_level = double(max(D(:)));
% my_map = jet(max_level);
% fig2 = figure;
%
% %intensity distribution - top 2/3 (nobkpt)
% subplot(3,1,1:2)
% hist(double(im(:)),max_level)
% axis([0 max_level 0 900])
% title('Distribution')
%
% %color scale - bottom 1/3 (nobkpt)
% subplot(3,1,3)
% imagesc(1:max_level)
% colormap(my_map)
% xlim([0 max_level])
% set(gca,'ytick',[])
% ylabel('Color Map')
% xlabel('Intensity')
% set(fig2,'position',[22 60 560 300],'render','zbuffer')
% set(fig1,'position',[601 68 392 314])
% figure(fig1)
% %% Segmentation
% %----------------------------------------------------------------------
%
% %ignore low levels (backround air, CSF & other soft? tissues)
% %using custom GUI tool to select best threshold level
% im = imrotate(squeeze(D(30,:,:)),90);
% figure(hSlico2)
%
% %remove some figures (no longer needed) - nobkpt
% if ishandle(fig1), delete(fig1), end
% if ishandle(fig2), delete(fig2), end
% doc graythresh
%
% %custom GUI tool (nobkpt)
% thresh_tool(im)
%
% %duplicate original data set for later reference (nobkpt)
% D1 = D;
%
% %apply some thresholding rules to ignore certain parts of data
% D(D<=40) = 0; %ignore low levels (CSF & air)
% D(D>=100) = 0; %ignore high levels (skull & other hard? tissues)
% D(:,:,1:60) = 0; %ignore spatially low positions (below brain mass)
% update_SliceOMatic(double(D),hSlico2)
%
% %erode away thick layer (dissolve thin surrounding tissues)
% blk = ones([3 7 7]);
% D = imerode(D,blk);
% update_SliceOMatic(double(D),hSlico2)
%
% %isolate brain mass (bwlabeln)
% doc bwlabel
% lev = graythresh(double(im)/max_level) * max_level;
% bw = (D>=lev);
% L = bwlabeln(bw);
%
% %connected region properties - how many, how big?
% doc regionprops
% stats = regionprops(L,'Area')
% A = [stats.Area];
% biggest = find(A==max(A))
%
% %remove smaller scraps
% D(L~=biggest) = 0;
% update_SliceOMatic(double(D),hSlico2)
%
% %grow back main region (brian mass) - nobkpt
% D = imdilate(D,blk);
% update_SliceOMatic(double(D),hSlico2)
%
% %separate white vs. gray matter
% im = imrotate(squeeze(D(30,:,:)),90);
% figure(hSlico2)
% lev2 = thresh_tool(im,'gray')
%
% %partition brain mass (nobkpt)
% lev2 = 67;
% L = zeros(size(D)); %0=outside brain (head/air)
% L(D<lev2 & D>0) = 2; %2=gray matter
% L(D>=lev2) = 3; %3=white matter
%
% %new Slice-O-Matic viewer (label matrix) - nobkpt
% sliceomatic(L)
% hSlico3 = gcf;
% daspect(1./voxel_size)
% set(hSlico3,'position',[455 63 560 420])
%
% %remove previous slicomatic viewer (nobkpt)
% if ishandle(hSlico2), delete(hSlico2), end
%
% %% Volumetric Measurements (voxel counting)
% %----------------------------------------------------------------------
%
% %total volume of brain (liters)
% brain_voxels = length(find(L(:)>1));
% brain_volume = brain_voxels*prod(voxel_size)/1e6
%
% %volume of gray matter (liters) - nobkpt
% gray_voxels = length(find(L(:)==2));
% gray_volume = gray_voxels*prod(voxel_size)/1e6
%
% %volume of white matter (liters) - nobkpt
% white_voxels = length(find(L(:)==3));
% white_volume = white_voxels*prod(voxel_size)/1e6
%
% %density calculations (volume ratios) - nobkpt
% gray_fraction = gray_volume/brain_volume
% white_fraction = white_volume/brain_volume
%
% return
%
% %% Separate head from background for visualization (advanced maneuver)
% %----------------------------------------------------------------------
%
% %duplicate data
% L1 = L;
%
% %exterior of head (connected inside through ears)
% %new Slice-O-Matic viewer (binary data) - nobkpt
% BW = (D1<lev1);
% %sliceomatic(double(BW))
% %hSlico3 = gcf;
% %daspect(1./voxel_size)
% %set(hSlico3,'position',[455 63 560 420])
%
% %melt away layer (separate small interior volumes) - nobkpt
% BW = imerode(BW,blk);
% %update_SliceOMatic(double(BW),hSlico3)
%
% %how many connected regions? (nobkpt)
% L = bwlabeln(BW);
% stats = regionprops(L,'Area')
%
% %which one biggest? (nobkpt)
% A = [stats.Area];
% biggest = find(A==max(A))
% BW(L~=biggest) = 0;
% %update_SliceOMatic(double(BW),hSlico3)
%
% %grow layer back (nobkpt)
% BW = imdilate(BW,blk);
%
% %label head voxels (different from air) - nobkpt
% L = L1;
% L(BW<1 & L1<2) = 1; %1=head (0=air, 2=gray, 3=white)
%
% %update final display (nobkpt)
% update_SliceOMatic(L,hSlico3)
%
% %% additional volumetric measurements
% %----------------------------------------------------------------------
%
% %volume ratio occupied by patient (nobkpt)
% scan_voxels = numel(L);
% head_voxels = length(find(L(:)>0));
% scan_density = head_voxels/scan_voxels
%
% %volume ratio of brain in head (nobkpt)
% brain_voxels = length(find(L(:)>1));
% brain_density = brain_voxels/head_voxels
| 15.zip | |
| File Size: | 2685 kb |
| File Type: | zip |